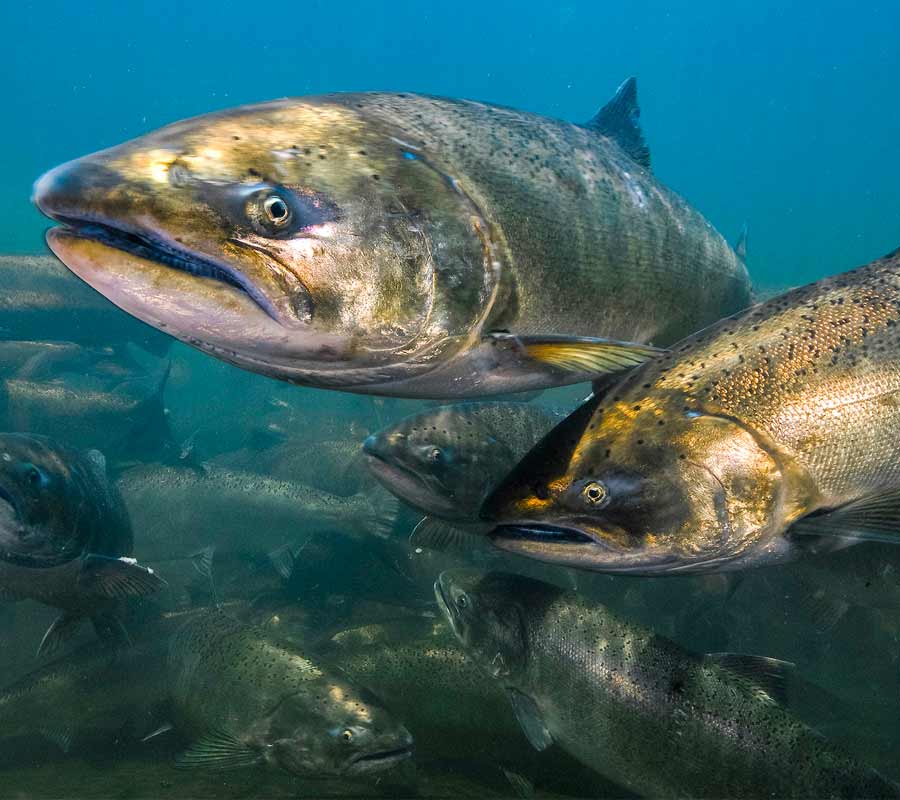
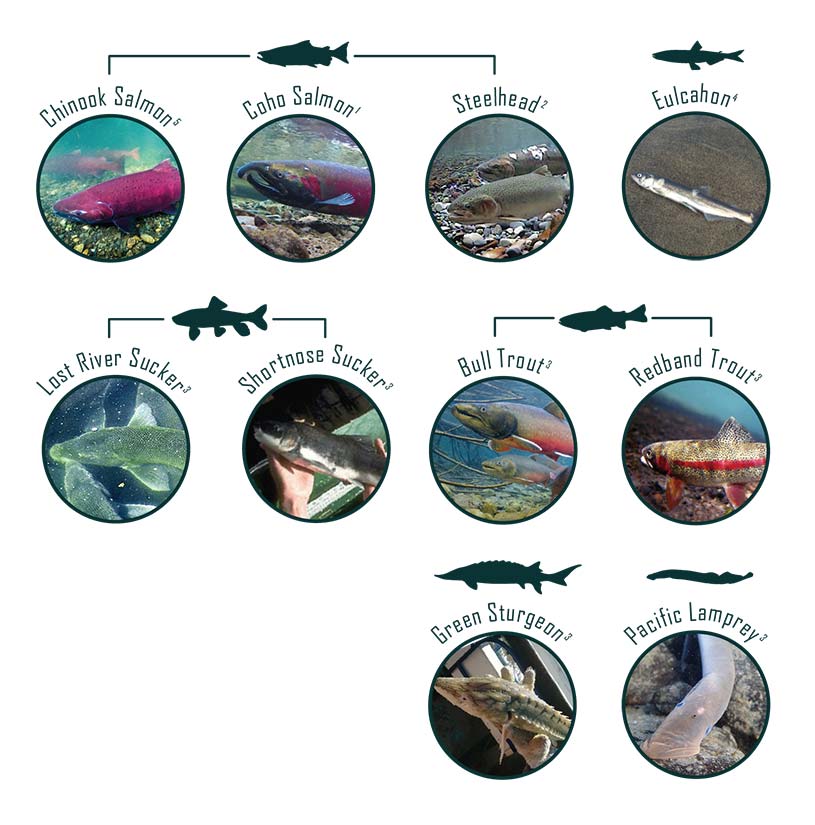
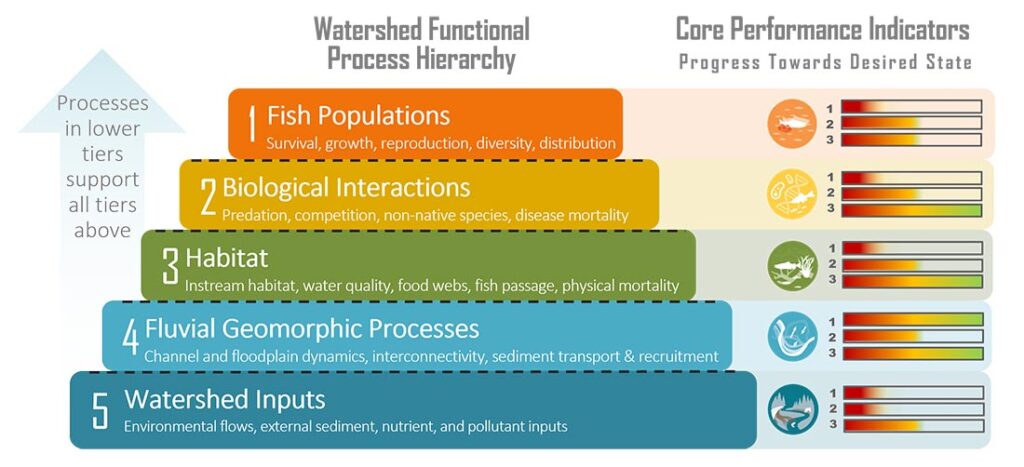
Potential Performance Indicator Suitability Thresholds
To assist with future monitoring and status reporting for the Klamath basin, ESSA researched potential suitability thresholds for candidate performance indicators and compiled available information